Antibody Characterization
In the development of antibody drugs, discovering the possible antibody candidates (hit) is only the first step. In this first step, the primary criteria that is typically used is binding affinity. Subsequently the antibody candidates need to be characterized and optimized for efficacy, safety and developability properties (lead). This characterization would entail in vitro and in vivo experiments which are time consuming and resource intensive.
In silico tools and capabilities can be leveraged to optimize this effort by evaluating the antibody candidates for their developability properties. While this is not an alternative to in vitro experiments, it helps in filtering out the antibodies with bad developability profile and focusing further in vitro experimentation on a select smaller set. AI/ML tools trained on developability property data can be used to characterize the antibodies and provide insights into whether they fall into the desired profile.
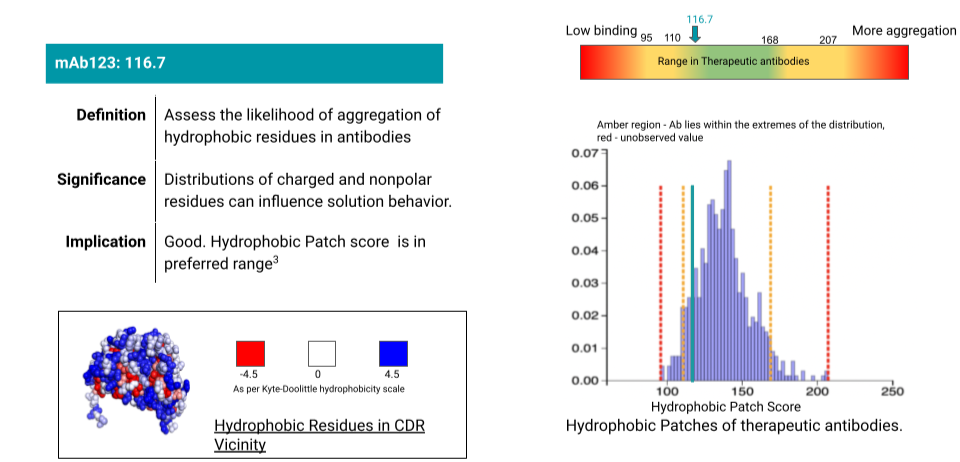
Aganitha provides an exhaustive developability property profile of the antibody including thermal stability, isoelectric point identification, hydrophobic patches, aggregation propensity, humanization score etc. The characterization report not only gives you a score for the antibody with respect to the property but also measures it against approved therapeutic antibodies. The report provides the implications of the score and its significance along with molecular images as appropriate (eg: hydrophobic patches). Below is a sample of the report for hydrophobic patches which are an indication of aggregation propensity of an antibody.
As part of your antibody drug discovery process, we recognize that there will be antibody candidates that exhibit good property profiles on some properties, and not so good property profiles on other properties. We can leverage our Generative AI capabilities to create new candidates to address the deficiencies.
We have the team and computational capabilities (research scientists, AI engineers, AI models, software tools) that can work with you to optimize a promising candidate (i.e create new nearby antibody sequences with better properties).
Download our Antibody Characterization report on select properties.
Learn more about our Antibody Engineering offerings
Antibody Virtual Screening
Antibody Lead Optimization
Generative AI based model to optimize properties such as humanization, aggregation optimization, stability improvement, etc. for increased precision.
Sequence Structure Co-Modeling
Explore the unexplored designable antibody space with generative AI models while considering sequence and structural co-modeling to design antibodies for a target antigen of interest.