mRNA Platform
In silico platform for biopharma scientists to design mRNA therapeutic candidates
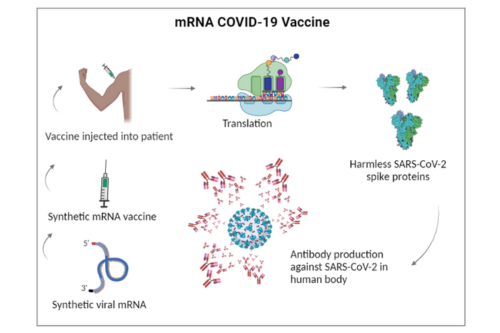
Success of mRNA vaccines against SARS-CoV-2 has demonstrated the great potential of mRNA based therapies. The potential of mRNA sequence to produce specific proteins using cells’ own translation machinery without causing any permanent alteration to the DNA sequence makes it an important modality.
Key challenges in mRNA design include:
- Selecting the optimal coding sequence of mRNA from a formidably large search space
- Multidimensionality of the problem space leading to increased expenses, manpower, and computation time
- Ensuring ex vivo and in vivo stability as well as translation efficiency
- Lack of customizable and scalable pipelines for complete mRNA characterisation and design
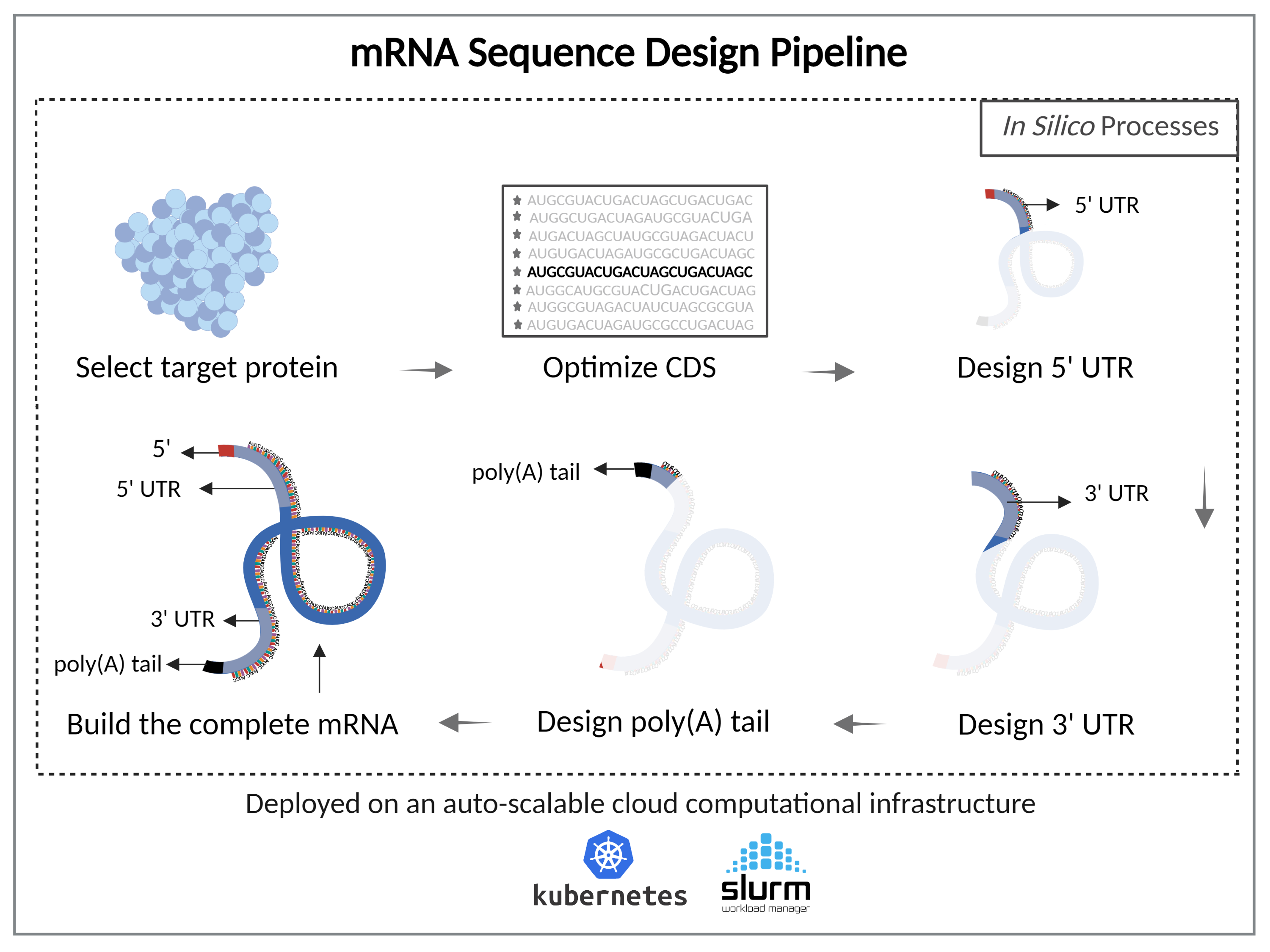
The platform consists of an integrated end-to-end pipeline for complete mRNA design. It offers researchers the ability to tailor an entire mRNA through:
- Target Protein Selection
- mRNA Coding Sequence Optimization
- Compatible non-coding Parts Design
The solution is deployed on a computational platform for high throughput screening and cloud-based Kubernetes environment with schedulers such as SLURM for auto-scaling and workload management.
Key steps of the mRNA pipeline include:
Optimize the coding region of the mRNA coding region (CDS)
Check the presence of Kozak consensus sequence in the designed mRNA
Highlights
Key components & strengths
An end-to-end solution for mRNA sequence design
Efficient algorithms for mRNA optimization
Customizable solution components
Cloud-based and on-demand computational infrastructure
Outcomes
Accelerate mRNA-based drug discovery and development
Reduced search space
Reduced cycle time
Faster selection
Scalable computation
Discover our offerings across the biopharma value chain
Our Solutions
Our Services
Offering services in computational sciences and technology to complement biopharma R&D